Many toxins synthesized by gram-negative bacteria are proteins harboring Repeat-in-ToXin (RTX) repeated motifs, typically composed of sequential glycine-rich nonapeptides of sequence GGXGXDXLX, where X stands for any amino acid and L is occasionally substituted by V, F or Y. One of the main features of these RTX motifs is their calcium-binding capacity. Adenylate cyclase toxin (CyaA), a protein of 1706 amino-acids, is one of the major virulence factors of Bordetella pertussis, the causative agent of whooping cough. CyaA is composed of three regions, the catalytic domain in its N-terminal part, followed by a hydrophobic region and a C-terminal RTX domain (RD) that contains tandem-repeated RTX motifs. CyaA RD (701 residues, 1006-1706 in native numbering) is organized in five successive blocks (I to V), each of 8 to 10 RTX motifs separated by linkers of variable length (from 23 to 49 amino-acids).
The apo-state (in the absence of calcium) of RD assumes an unfolded conformation, which exhibits the biophysical features of the so-called intrinsically disordered proteins. RD undergoes a calcium-induced compaction leading to the holo-state, which is characterized by stable secondary and tertiary structures (see J. Biol. Chem., Jan 2009; 284: 1781 - 1789). This application note presents the characterization of the two states of RD by Size Exclusion Chromatography (SEC) coupled with the Triple Detection Array (TDA). Using SEC-TDA, the absolute molecular mass is measured as well as the intrinsic viscosity. Since the intrinsic viscosity (IV) is dependent on the density, hydration and overall shape of the polypeptide, a major change in IV is expected between the apo- and holo-RD conformations.
The hydrodynamic behavior of apo- and holo-RD was explored using a Viscotek Triple Detector Array (TDA302) system, coupled to a GPCmax chromatographic system equipped with a Superdex 200 column (GE Healthcare) for the efficient separation of RD. The elution buffer used for apo-RD was a solution of 5 mM Hepes, 100 mM NaCl at pH 7.3. For holo-RD, 2 mM CaCl2 was added to the buffer. According to the general procedure recommended by Viscotek, all the solutions were filtered through a 0.2 µm filter and allowed to equilibrate at 10 °C. SEC elution was done at 10 °C and the measurements were made at 20 °C. Bovine Serum Albumin (BSA) injections were used to determine the inter-detector delay, the calibration constants as well as the band broadening correction factor.
The OmniSEC software was used to control the GPCmax integrated degasser, pump and autosampler system. The same software was used to acquire the signals from the 5 detectors of the TDA system: a UV-Visible spectrophotometer, a 7° Low Angle Light Scattering (LALS) and a 90° Right Angle Light Scattering (RALS) detectors measuring the static light scattering intensity, a differential refractometer (RI) and a differential pressure (DP) Wheatstone bridge viscometer, all in series. OmniSEC was used to calculate the absolute molecular mass (MM) as well as the intrinsic viscosity (IV) of the protein from the above-mentioned measurements (RI, UV, LALS, RALS and DP).
Injections at various concentrations of RD in both buffers (5 mM Hepes, 100 mM NaCl ± 2 mM CaCl2 at pH 7.3) were performed to determine experimentally the refractive index increment, dn/dc, of the apo- and holo-states. We found that, in both cases, the dn/dc values were identical and equal to 0.184 cm3/g (Table 1).
| Parameter | Units | apo-RD | holo-RD |
|---|---|---|---|
| Elution volume | mL | 10.4 | 13.8 |
| dn/dc | cm3/g | 0.184 | 0.184 |
| Intrinsic viscosity | dL/g | 0.35 | 0.055 |
| Molecular mass | kg/mol | 73.6 | 73.2 |
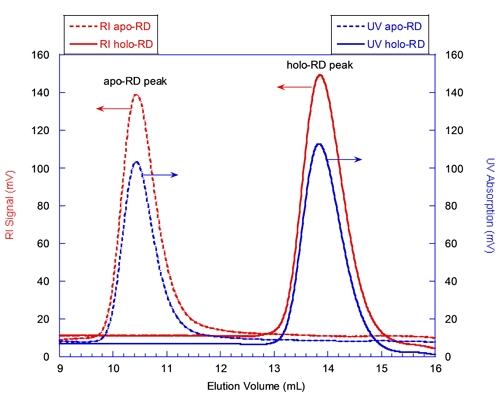
The solid lines in figure 1 correspond to the elution of holo-RD while the dashed lines correspond to that of apo-RD. The red and blue curves correspond respectively to the RI and UV detectors, whose signals are proportional to the sample concentration. Both forms of RD eluted as a single narrow peak. The elution volume of holo-RD was 13.8 mL. This elution volume is close to that expected for a globular and folded protein of 73 kDa, which would elute at 15 mL. In marked contrast, the elution volume of apo-RD was 10.4 mL, a value corresponding to a folded, globular protein of approximately 600 kDa. Hence, conventional SEC measurements (which assume that the retention volume is directly related to the MM) would have suggested that the MM of both forms of RD were largely different, apo-RD behaving as a multimer of 600 kDa, and holo-RD as a monomer.
|
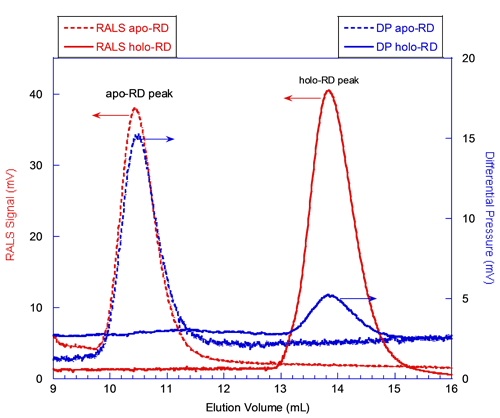
In figure 2, the red curves correspond to the signals from the RALS detector. The static light scattering signal is proportional to the product of the molecular mass and the concentration of the protein deduced from RI or UV detectors (International Laboratory News, April 2006, M. Haney, p.12, 14). Hence, the absolute molecular mass is calculated for each species, independently from the elution volume. At first glance, the comparison of the RALS chromatograms of apo-RD and holo-RD (Figure 2) shows that peak heights of both proteins are comparable. This indicates that apo-RD and holo-RD have similar molecular masses. Calculations with the OmniSEC software confirmed this observation, providing MM of 73.6 and 73.2 kDa for apo- and holo-RD, respectively. Hence, both forms of RD are in fact monomers (Table 1).
|
The blue curves in figure 2 correspond to the signals from the Differential Pressure (DP) transducer of the viscometer. The DP signal for apo-RD is stronger than the one for holo-RD (Figure 2). Indeed, the OmniSEC software calculated an intrinsic viscosity (Table 1) of 0.35 and 0.055 dL/g for apo- and holo-RD, respectively. The intrinsic viscosity is related to the hydration and to the overall shape (spherical or elongated shape), and is inversely proportional to the density of the polypeptide. The fact that the intrinsic viscosity of apo-RD is significantly higher than that of holo-RD suggests that RD might adopt a hydrated and/or elongated conformation in the absence of calcium. This huge difference of intrinsic viscosity reflects the dramatic hydrodynamic and conformational changes induced by the binding of calcium to RD.
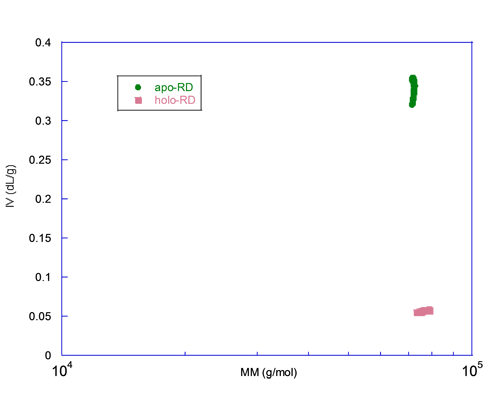
A graph of the intrinsic viscosity (IV) versus the molecular mass (MM) measured for each slice centered on the peak is plotted in figure 3. This figure illustrates that both forms have the same MM, here again ruling out aggregation or oligomerization, but the IV differs by almost an order of magnitude. Since the IV is inversely proportional to the density and related to the shape, apo-RD is expected to be highly hydrated and/or to adopt a rod-like shape. These hydrodynamic parameters, hydration and shape of both apo and holo states, are characterized in detail elsewhere (J. Biol. Chem., Jan 2009; 284: 1781 - 1789). In conjunction with the hydrodynamic radii (Rh) of apo-RD and holo-RD obtained either by analytical ultracentrifugation (AUC) or by Dynamic Light Scattering (DLS), we showed that (i) apo-RD is highly hydrated and that (ii) the binding of calcium induces a strong dehydration and an elongation of RD.
|
In this note, we show that size exclusion chromatography followed by triple detection array (SEC-TDA) is an appropriate method to investigate the hydrodynamic and conformational changes of polypeptides. Conventional SEC, based on the retention volumes of folded and globular standard proteins, could not distinguish between oligomerization, hydration or shape changes. Here, SEC-TDA clearly indicates that both apo- and holo-states of RD are monomeric. Finally, we show that intrinsic viscosity is the only hydrodynamic parameter that is significantly affected by calcium binding. These results indicate that calcium binding induces important hydrodynamic changes, such as dehydration and molecular shape changes.
This application note is based on the paper published in J. Biol. Chem., Jan 2009; 284: 1781 - 1789.
Alexandre Chenal, Bertrand Raynal, Patrick England and Daniel Ladant, Institut Pasteur, Paris, France.